Biomolecular Networks Laboratories
Laboratory of Chromosome Biology
 Prof. FUKAGAWA Tatsuo
Prof. FUKAGAWA Tatsuo
Keywords:
Chromosome, Kinetochore, Centromere, Chromosome segregation
How are chromosomes segregated into daughter cells?
The kinetochore plays a fundamental role in accurate chromosome segregation during mitosis and meiosis in eukaryotes. Although chromosome segregation errors cause genetic diseases, including some cancers, the mechanisms of how kinetochores interact with microtubules of the spindle apparatus during cell division are not fully understood. To understand the molecular mechanisms of chromosome segregation, we are focusing on the kinetochore and are trying to address how the kinetochore is specified and assembled on centromere chromatin. In addition, we are analyzing how the kinetochore correctly attaches to microtubules. We are utilizing molecular biology, cell biology, biochemistry, structural biology, and genome engineering to clarify kinetochore structure and function.
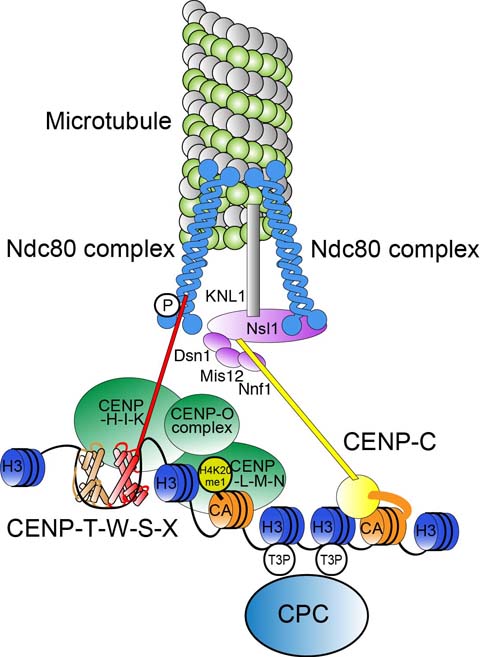
Molecular model of the kinetochore structure. We would like to address what the structure is and how the structure is established and functions.
Members
| FUKAGAWA Tatsuo (Professor) | fukagawa.tatsuo.fbs[at]osaka-u.ac.jp |
|---|---|
| HORI Tetsuya (Associate Professor) | t.hori.fbs[at]osaka-u.ac.jp |
| ASAKAWA Haruhiko (Associate Professor) | asakawa.haruhiko.fbs[at]osaka-u.ac.jp |
| HIRANO Yasuhiro (Specially Appointed Lecturer) | y.hirano.fbs[at]osaka-u.ac.jp |
| ARIYOSHI Mariko (Specially Appointed Assistant Professor) | ariyoshi.mariko.fbs[at]osaka-u.ac.jp |
| SRIDHAR Shreyas (Specially Appointed Assistant Professor) | sridhar.shreyas.fbs[at]osaka-u.ac.jp |
| TAKENOSHITA Yusuke (Specially Appointed Researcher) | takenoshita.yusuke.fbs[at]osaka-u.ac.jp |
| MAHANA Yutaka (Specially Appointed Researcher) | mahana.yutaka.fbs[at]osaka-u.ac.jp |
| MAKINO Fumiaki (Visiting Researcher) | |
| KUBOTA Yoshino (Technical Staff) | |
| FUKUOKA Reika (Technical Staff) |
You could probably reach more information of individual researchers by Research Map and researcher's search of Osaka-U.
- ※Change [at] to @
Q&A
- What is your hot research topic?
- We are focusing on the question "how is the kinetochore structure is organized?" Since identification of most kinetochore components, many groups are trying to address this question using in vitro reconstitution with recombinant components. However, the kinetochore structure is dynamically changed during progression of the cell cycle, and we would like to clarify how such dynamic changes occur.
- What is your breakthrough or research progress in the last 5 years?
- The most important role for the kinetochore is to bridge a connection between chromatin and spindle microtubules. Therefore, it is critical to clarify how microtubule binding proteins are recruited into the kinetochore. While many groups proposed that CENP-C is critical for the recruitment of microtubule binding proteins into the kinetochores, we demonstrated that CENP-T is much more critical for this event. This research result broke the mold. Furthermore, we revealed how CENP-C associates with chromatin, based on structure analyses.
- What kind of background do your lab members have?
- Lab members have backgrounds in science, agriculture, medicine or pharmaceutical sciences. As various techniques are needed to conduct centromere/kinetochore research, we are seeking people from various fields.
- Do you collaborate with other institutions and universities?
- We are collaborating with scientists from the University of Tokyo, Tokyo tech., the National Institute of Genetics, MIT, the University of Edinburgh, and the Curie Institute.
- What kind of careers do your Lab's alumni go on to?
- Many of them get research jobs in academia and the business sector. However, some people also work in non-scientific fields.
- How do you develop your research?
- We mainly used cultured-cells for our centromere/kinetochore studies. However, we would like to study on various mechanisms for chromosomes segregation in organismal context in the future.
Research Highlights
Publications (Research Articles, Reviews, Books)
2025
Exploring the cell nucleus: From chromosome structure to single-cell omics
Curr. Opin. Cell Biol. 95:102530. 2025 (PMID:40381430 DOI:10.1016/j.ceb.2025.102530)
Meiosis: When centromeres choose compromise over conflict
Curr. Biol. 35(5):R196-R198. 2025 (PMID:40068619 DOI:10.1016/j.cub.2025.01.059)
Interaction mapping between nucleoporins in the fission yeast Schizosaccharomyces pombe using mass-spectrometry
J. Biochem. 177(4):273-286. 2025 (PMID:39727334 DOI:10.1093/jb/mvae095)
2024
Molecular details and phosphoregulation of the CENP-T-Mis12 complex interaction during mitosis in DT40 cells
iScience 27(12):111295. 2024 (PMID:39628583 DOI:10.1016/j.isci.2024.111295)
CENP-C-Mis12 complex establishes a regulatory loop through Aurora B for chromosome segregation
Life Sci. Alliance 8(1):e202402927. 2024 (PMID:39433344 DOI:10.26508/lsa.202402927)
Disordered region of nuclear membrane protein Bqt4 recruits phosphatidic acid to the nuclear envelope to maintain its structural integrity
J. Biol. Chem. 300(7):107430. 2024 (PMID:38825008 DOI:10.1016/j.jbc.2024.107430)
Vertebrate centromeres in mitosis are functionally bipartite structures stabilized by cohesin
Cell 187(12):3006-3023.e26. 2024 (PMID:38744280 DOI:10.1016/j.cell.2024.04.014)
Artificial tethering of constitutive centromere-associated network proteins induces CENP-A deposition without Knl2 in DT40 cells
J. Cell Sci. 137(2): cs261639. 2024 (PMID:38319136 DOI:10.1242/jcs.261639)
2023
An updated view of the kinetochore architecture
Trends Genet. 39(12):941-953. 2023 (PMID:37775394 DOI:10.1016/j.tig.2023.09.003)
CENP-A and CENP-B collaborate to create an open centromeric chromatin state
Nature Communu. 14(1):8227. 2023 (PMID:38086807 DOI:10.1038/s41467-023-43739-5)
Mitotic perturbation is a key mechanism of action of decitabine in myeloid tumor treatment
Cell Rep. 42(9):113098. 2023 (PMID:37714156 DOI:10.1016/j.celrep.2023.113098)
A ubiquitin-proteasome pathway degrades the inner nuclear membrane protein Bqt4 to maintain nuclear membrane homeostasis
J. Cell Sci. 136(19):jcs260930. 2023 (PMID:37694715 DOI:10.1242/jcs.260930)
Centromere/kinetochore is assembled through CENP-C oligomerization
Mol. Cell 83(13):2188-2205.e13. 2023 (PMID:37295434 DOI:10.1016/j.molcel.2023.05.023)
Ceramide synthase homolog Tlc4 maintains nuclear envelope integrity via its Golgi translocation
J. Cell Sci. 136(10):jcs260923. 2023 (PMID:37078207 DOI:10.1242/jcs.260923)
Evolutionary analysis of a complete chicken genome
Proc. Natl. Acad. Sci. USA 120(8):e2216641120. 2023 (PMID:36780517 DOI:10.1073/pnas.2216641120)
The cryo-EM structure of the CENP-A nucleosome in complex with ggKNL2
EMBO J. 42(6):e111965. 2023 (PMID:36744604 DOI:10.15252/embj.2022111965)
Transgenerational epigenetic control of constitutive heterochromatin, transposons, and centromeres
Curr. Opin. Genet. Dev. 78:102021. 2023 (PMID:36716679 DOI:10.1016/j.gde.2023.102021)
2022
Kinetochore Architecture Employs Diverse Linker Strategies Across Evolution
Front. Cell Dev. Biol. 10:862637. 2022 (PMID:35800888 DOI:10.3389/fcell.2022.862637)
Recruitment of two Ndc80 complexes via the CENP-T pathway is sufficient for kinetochore functions
Nature Commun. 13(1):851. 2022 (PMID:35165266 DOI:10.1038/s41467-022-28403-8)
Transfected plasmid DNA is incorporated into the nucleus via nuclear envelope reformation at telophase
Commun. Biol. 5(1):78. 2022 (PMID:35058555 DOI:10.1038/s42003-022-03021-8)
Mobility of kinetochore proteins measured by FRAP analysis in living cells
Chromosome Res. 30:43-57. 2022 (PMID:34997387 DOI:10.1007/s10577-021-09678-x)
A Simple Method that Combines CRISPR and AID to Quickly Generate Conditional Knockouts for Essential Genes in Various Vertebrate Cell Lines
Methods Mol. Biol. 2377:109-122. 2022 (PMID:34709613 DOI:10.1007/978-1-0716-1720-5_6)
2021
Bub1 and CENP-U redundantly recruit Plk1 to stabilize kinetochore-microtubule attachments and ensure accurate chromosome segregation
Cell Rep. 36(12):109740. 2021 (PMID:34551298 DOI:10.1016/j.celrep.2021.109740)
A Simple Method to Generate Super-sensitive AID (ssAID)-based Conditional Knockouts using CRISPR-based Gene Knockout in Various Vertebrate Cell Lines
Bio Protocol e11(14):e4092. 2021 (PMID:34395730 DOI:10.21769/BioProtoc.4092)
Kinetochore stretching-mediated rapid silencing of the spindle-assembly checkpoint required for failsafe chromosome segregation
Curr. Biol. 31(8):1581-1591.e3. 2021 (PMID:33651990 DOI:10.1016/j.cub.2021.01.062)
CENP-C Phosphorylation by CDK1 in vitro
Bio Protocol 11(1):e3879. 2021 (PMID:33732767 DOI:10.21769/BioProtoc.3879)
Bridgin connects the outer kinetochore to centromeric chromatin
Nature Commun. 12(1):146. 2021 (PMID:33420015 DOI:10.1038/s41467-020-20161-9)
Cryo-EM structure of the CENP-A nucleosome in complex with phosphorylated CENP-C
EMBO J. 40(5):e105671. 2021 (PMID:33463726 DOI:10.15252/embj.2020105671)
2020
A super-sensitive auxin-inducible degron system with an engineered auxin-TIR1 pair.
Nucl. Acids. Res. 48(18):e108. 2020 (PMID:32941625 DOI:10.1093/nar/gkaa748)
H3K9me3 Maintenance on a Human Artificial Chromosome Is Required for Segregation but Not Centromere Epigenetic Memory.
J. Cell Sci. 133(14):jcs242610. 2020 (PMID:32576667 DOI:10.1242/jcs.242610)
CENP-50 Is Required for Papilloma Development in the Two-Stage Skin Carcinogenesis Model.
Cancer Sci. 111(8):2850-60 2020 (PMID:32535988 DOI:10.1111/cas.14533)
Dynamics of kinetochore structure and its regulations during mitotic progression.
Cell. Mol. Life Sci. 77(15):2981-2995. 2020 (PMID:32052088 DOI:10.1007/s00018-020-03472-4)
Artificial generation of centromeres and kinetochores to understand their structure and function.
Exp. Cell Res. 389(2):111898 2020 (PMID:32035949 DOI:10.1016/j.yexcr.2020.111898)
2019
CDK1-mediated CENP-C phosphorylation modulates CENP-A binding and mitotic kinetochore localization
J. Cell Biol. 218(12):4042-4062 2019 (PMID:31676716 DOI:10.1083/jcb.201907006)
Cryo-EM Structures of Centromeric Tri-nucleosomes Containing a Central CENP-A Nucleosome
Structure 28(1):44-53.e4 2019 (PMID:31711756 DOI:10.1016/j.str.2019.10.016)
Centromere maintenance during DNA replication
Nat. Cell Biol. 21(6):669-671 2019 (PMID:31160706 DOI:10.1038/s41556-019-0335-0)
Where is the right path heading from the centromere to spindle microtubules?
Cell Cycle 18(11):1199-1211 2019 (PMID:31075048 DOI:10.1080/15384101.2019.1617008)
Live imaging of marked chromosome regions reveals their dynamic resolution and compaction in mitosis.
J. Cell Biol. 218(5):1531-1552 2019 (PMID:30858191 DOI:10.1083/jcb.201807125)
The CENP-A centromere targeting domain facilitates H4K20 monomethylation in the nucleosome by structural polymorphism.
Nat. Commun. 10(1):576 2019 (PMID:30718488 DOI:10.1038/s41467-019-08314-x)
2018
Multiple phosphorylations control recruitment of the KMN-network onto kinetochores
Nat. Cell Biol. 20(12):1378-1388 2018 (PMID:30420662 DOI:10.1038/s41556-018-0230-0)
3D genomic architecture reveals that neocentromeres associate with heterochromatin regions
J. Cell Biol. 218(1):134-149 2018 (PMID:30396998 DOI:10.1083/jcb.201805003)
Cooperative Domain Formation by Homologous Motifs in HOIL-1L and SHARPIN Plays A Crucial Role in LUBAC Stabilization.
Cell Reports 23(4):1192-1204 2018 (PMID:29694895 DOI:10.1016/j.celrep.2018.03.112)
Kinetochore assembly and disassembly during mitotic entry and exit.
Curr. Opin. Cell Biol. 52:73-81 2018 (PMID:29477052 DOI:10.1016/j.ceb.2018.02.005)
Identification of PNG kinase substrates uncovers interactions with the translational repressor TRAL in the oocyte-to-embryo transition.
eLife 7:e33150 2018 (PMID:29480805 DOI:10.7554/eLife.33150)
Site-Specific Cleavage by Topoisomerase 2: A Mark of the Core Centromere.
Int. J. Mol. Sci. 19(2):E534 2018 (PMID:29439406 DOI:10.3390/ijms19020534)
2017
Structure of the Dnmt1 Reader Module Complexed with a Unique Two-Mono-Ubiquitin Mark on Histone H3 Reveals the Basis for DNA Methylation Maintenance.
Mol. Cell 68(2):350-360.e7 2017 (PMID:29053958 DOI:10.1016/j.molcel.2017.09.037)
Critical Foundation of the Kinetochore: The Constitutive Centromere-Associated Network (CCAN).
Prog Mol Subcell Biol. 56:29-57 2017 (PMID:28840232 DOI:10.1007/978-3-319-58592-5_2)
CENP-R acts bilaterally as a tumor suppressor and as an oncogene in the two-stage skin carcinogenesis model.
Cancer Sci. 108(11):2142-2148 2017 (PMID:28795467 DOI:10.1111/cas.13348)
An efficient method to generate conditional knockout cell lines for essential genes by combination of auxin-inducible degron tag and CRISPR/Cas9.
Chromosome Res. 25(3-4):253-260 2017 (PMID:28589221 DOI:10.1007/s10577-017-9559-7)
Assembly and stoichiometry of the core structure of the bacterial flagellar type III export gate complex.
PLoS. Biol. 15(8):e2002281 2017 (PMID:28771466 DOI:10.1371/journal.pbio.2002281)
Straight and rigid flagellar hook made by insertion of the FlgG specific sequence into FlgE.
Sci Rep 7:46723 2017 (PMID:28429800 DOI:10.1038/srep46723)
Association of M18BP1/KNL2 with CENP-A nucleosome is essential for centromere formation in non-mammalian vertebrates
Dev. Cell 42(2):181-189.e3 2017 (PMID:28743004 DOI:10.1016/j.devcel.2017.06.019)
Critical histone post-translational modifications for centromere function and propagation
Cell Cycle 16(13):1259-1265 2017 (PMID:28840232 DOI:10.1080/15384101.2017.1325044)
Control of PNG kinase, a key regulator of mRNA translation, is coupled to meiosis completion at egg activation
eLife 6:e22219 2017 (PMID:28555567 DOI:10.7554/eLife.22219)
Acute inactivation of the replicative helicase in human cells triggers MCM8-9-dependent DNA synthesis
Genes Dev. 31(8):816-829 2017 (PMID:28487407 DOI:10.1101/gad.297663.117)
Stepwise unfolding supports a subunit model for vertebrate kinetochores
Proc. Natl. Acad. Sci. U. S. A. 114(12):3133-3138 2017 (PMID:28265097 DOI:10.1073/pnas.1614145114)
Constitutive centromere-associated network controls centromere drift in vertebrate cells
J. Cell Biol. 216(1):101-113 2017 (PMID:27940888 DOI:10.1083/jcb.201605001)
2016
The architecture of the cytoplasmic region of type III secretion systems.
Sci Rep 6:33341 2016 (PMID:27686865 DOI:10.1038/srep33341)
Acetylation of histone H4 Lysine 5 and 12 is required for CENP-A deposition into centromeres
Nat. Commun. 7:13465 2016 (PMID:27811920 DOI:10.1038/ncomms13465)
Chromatin determinants of the inner-centromere rely on replication factors with functions that impart cohesion.
Oncotarget 7(42):67934-67947 2016 (PMID:27636994 DOI:10.18632/oncotarget.11982)
Kinetochore assembly and function through the cell cycle.
Chromosoma 125(4):645-59 2016 (PMID:27376724 DOI:10.1007/s00412-016-0608-3)
Chicken rRNA Gene Cluster Structure.
PLoS One 11(6):e0157464 2016 (PMID:27299357 DOI:10.1371/journal.pone.0157464)
Biochemical and Structural Analysis of Kinetochore Histone-Fold Complexes.
Methods Mol Biol. 1413:135-46 2016 (PMID:27193847 DOI:10.1007/978-1-4939-3542-0_9)
Centromeric chromatin and kinetochore assembly in vertebrate cells.
F. Hanaoka, and K. Sugasawa (eds.) DNA replication, recombination, and repair: Molecular Mechanisms and pathology 365-387 2016 ( DOI:10.1007/978-4-431-55873-6_14)
Genetic complementation analysis showed distinct contributions of the N-terminal tail of H2A.Z to epigenetic regulations.
Genes Cells 21(2):122-35 2016 (PMID:26833946 DOI:10.1111/gtc.12327)
Auxin/AID versus conventional knockouts: distinguishing the roles of CENP-T/W in mitotic kinetochore assembly and stability.
Open Biol 6(1):150230 2016 (PMID:26791246 DOI:10.1098/rsob.150230)
Our ideal candidate (as a graduate student)
We are looking for a highly motivated person to work on our research topics as our lab member. Our lab welcomes the person who loves taking care of creatures, hand working and handcraft too. Any kind of background (such as your expertise or major) is available.
Contact
Laboratory of Chromosome Biology, Graduate School of Frontier Biosciences, Osaka University,
1-3 Yamadaoka, Suita, Osaka 565-0871 Japan.
TEL:+81-6-6879-4428
E-mail: fukagawa.tatsuo.fbs[at]osaka-u.ac.jp (Prof. Tatsuo Fukagawa)
- ※Change [at] to @
